SECOND ANNUAL
Crops in silico
SYMPOSIUM AND WORKSHOP
June 26-28th 2017
University of Oxford, UK
Select Presentations for Download
- Education and Community Building (in Astrophysics)- Gabrielle Allen, University of Illinois
- Semantic Annotation of Biosimulation Models for Model Reuse and Composition – Brian Carlson, University of Michigan
- Models for Model Integration – Daniel S. Katz, University of Illinois
- Signal Integration and the Bud Activation Switch – Ottoline Leyser, Cambridge University
- Templates, Anchors, and Matryoshkas: A Vision for Crop Systems Biology – Eberhard O. Voit, Georgia Institute of Technology
Meeting Recap
The second annual Crops in silico Symposium and Workshop was hosted at St. Catherine’s College, University of Oxford, on June 26-28, 2017. The goals of this Symposium and Workshop were:
- Briefly introduce the Crops in silico concept as an integrative and multi-scale modeling platform.
- Learn from plant and computer scientists who have progressed plant modeling or tools to enhance modeling in different domains.
- Discuss where sustainable and practical collaborations may be valuable and achievable in bridging layers of organization.
- Map the next steps for the community
57 registrants from 23 institutions and companies and six countries convened for the Symposium.
The event kicked off June 26 with Keynote Speaker Eberhard Voit, Professor and GRA Eminent Scholar and Flanagan Chair in Biological Systems at the Georgia Institute of Technology. His talk addressed the difficulties of modelling complex biological systems and how those difficulties can be overcome by dividing a complex modeling task into a coarse, framework model and more detailed, finer-grained models of subsystems.
The meeting included five presentations on recent developments in plant modelling at different levels of biological organization, three presentations on recent developments in model integration, four presentations on users and usability of models, and one presentation outlining the steps to the successful deployment of a community driven software platform to advance and support research in astrophysics.
The overall goal of the workshop was to identify ways to build the Cis community. The first part of the workshop focused on identifying ways to establish successful research collaborations. Next, attendees brainstormed ideas of how to build a Crops in silico research community. The second portion of the workshop was focused on fostering new research collaborations.
- 2nd annual Symposium and Workshop participants.
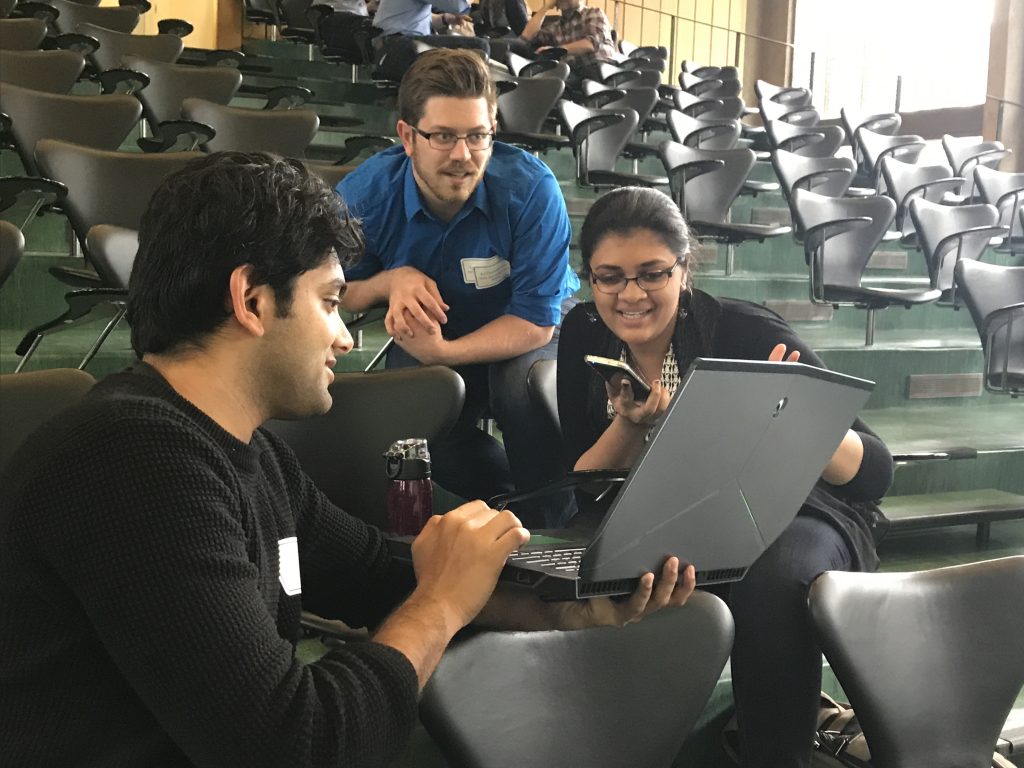
- Speaker discussion session
- Workshop: Building the Crops in silico community.
- Workshop: fostering new research collaborations
- Workshop: fostering new research collaborations
- Interactive poster session
Who should attend?
Join experts in experimentation, agronomy, physiology, plant development, phenotyping, as well as experts in computational modeling, software development, and data visualization.
agenda
Download the program.
5 – 7 pm
Keynote talk – Templates, Anchors and Matryoshkas: A vision for crop systems biology / Everhard Voit, Georgia Tech University
This talk is free and open to the public.
Crops have been the subject of unending attempts for improvement since the dawn of humanity, first through trial and error and later through selective breeding. Indeed, comparing modern crops with their early predecessors, one can only be astounded by the success of these methods. Unfortunately, both trial and error and selective breeding require much time, thus raising the question of whether the process can be sped up through rational crop modifications. The challenge of such an approach is the enormous complexity of plants. The Norway Spruce has close to 60,000 genes, and metabolites within the plant kingdom are estimated to range collectively between 200,000 and 1 million. Confounding these numbers are other features of complexity, including simultaneous operation at multiple scales, counterintuitive nonlinearities, synergistic and antagonistic effects, and unpredictable threshold effects. The convolution of these features cannot be grasped reliably by the unaided human mind and suggests the employment of methods of computational systems biology. In this presentation, I will sketch the contextual landscape in which computational systems biology might help advance the goals of crop science and propose a modular strategy for realizing the development of effective models.
7 pm
Conference dinner
Breakfast
Registration
9:30 am – 1:00 pm
Session 1: Recent developments in plant modelling
Cassava source-sink – Expression of genes involved in central carbon metabolism / Frank Ludewig, Universität Erlangen-Nürnberg
Constraints-based modelling of leaf metabolic networks / Lee Sweetlove, University of Oxford
Signal integration in the control of shoot branching / Ottoline Leyser, Cambridge University
Organ scale models: lessons from the hidden half / Malcolm Bennett, University of Nottingham
Crop modelling to underpin food security / Matt Reynolds, CGIAR
1:00 – 2:00 pm
Lunch
2:00 – 5:00 pm
Session 2: Recent developments in model integration
Models for model interoperability / Daniel S. Katz, University of Illinois
Integrative Dynamic Modeling Using Diverse Biological Datasets / Cranos M. Williams, North Carolina State University
A Computational Framework for Connecting Models Across Languages and Scales / Meagan Lang, National Center for Supercomputing Applications
Workshop: Identifying gaps, challenges, and potential solutions
7 pm
Conference dinner
8 – 9 am
Breakfast
9 – 1 pm
Session 3: Users and usability
Four Levels of Visualization / Min Chen, University of Oxford
Semantic Annotation of Biosimulation Models for Model Reuse and Composition / Brian Carlson, SemGen
Education Using Crops in silico / Gabrielle Allen, University of Illinois
Scaling from mechanistic models to crop stands and beyond / Steve Long, University of Illinois
Workshop: Identifying users and opportunities to enhance usability
1 – 2 pm
Lunch
Session 4: Next steps for Crops in silico
Workshop: Identifying crop improvement goals
Workshop: Developing future collaborations
Speakers

Eberhard Voit, Keynote Speaker
Professor and GRA Eminent Scholar, Flanagan Chair in Biological Systems, Georgia Institute of Technology

Gabrielle D. Allen
Professor of Astronomy and Associate Dean for Research and Research Education, University of Illinois

Malcolm Bennett
Professor of Plant Sciences, The University of Nottingham

Brian Carlson
Research Assistant Professor, Molecular & Integrative Physiology, University of Michigan

Min Chen
Professor of Scientific Visualisation, Oxford University

Daniel S. Katz
Assistant Director for Scientific Software and Applications, National Center for Supercomputing Applications & Research Associate Professor, Electrical and Computer Engineering & Computer Science, University of Illinois

Meagan Lang
Postdoctoral Research Associate in the National Center for Supercomputing Applications at the University of Illinois

Ottoline Leyser
Director of the the Sainsbury Laboratory, Cambridge University

Steve Long
Gutgsell Endowed Professor, Department of Crop Sciences and Plant Biology, University of Illinois
Newton Abraham Visiting Professor,Department of Plant Science, Oxford University

Frank Ludewig
Scientific coordinator of the Cassava Source-Sink project, Universität Erlangen-Nürnberg

Matt Reynolds
Wheat Physiologist, CGIAR and International Maize and Wheat Improvement Center

Cranos M. Williams
Director of the EnBiSys Research Laboratory and Assistant Professor of Electrical and Computer Engineering at North Carolina State University